Welcome to the inaugural Friday links post. We’ll be using these posts to share interesting articles stumbled across by Unzipped members during the week.
We’re still tweaking the format, but the basic idea will be a brief paragraph of commentary followed by the initials of the person who wrote it.
Dan Koboldt reviews a recent paper reporting the use of whole-genome sequencing to find the mutation responsible for a severe genetic disease. Interestingly, in this case the disease was undiagnosed, and the causal variant was used to produce a diagnosis of sitosterolemia; more interestingly, this diagnosis had already been ruled out by another test, that was shown to be a false negative. [DM]
 ScienceNews reports that researchers from the University of Copenhagen have got permission to sequence the genome of Sitting Bull, the native American war chief that led the battle of Little Bighorn. I don’t know exactly what they intend to learn from the genome scientifically, but it seems like this might serve primarily as a monument to a major figure in native American resistance. So the question I have is this: how can we go from a genome sequence (which is generally just a text file on a computer) to a public rememberance, something akin to the 1989 postage stamp shown to the left? [LJ]
ScienceNews reports that researchers from the University of Copenhagen have got permission to sequence the genome of Sitting Bull, the native American war chief that led the battle of Little Bighorn. I don’t know exactly what they intend to learn from the genome scientifically, but it seems like this might serve primarily as a monument to a major figure in native American resistance. So the question I have is this: how can we go from a genome sequence (which is generally just a text file on a computer) to a public rememberance, something akin to the 1989 postage stamp shown to the left? [LJ]
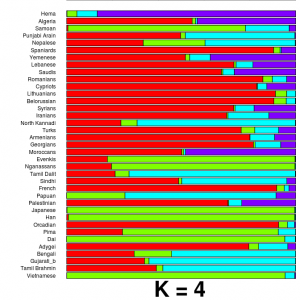
Two papers in the current issue of Nature Genetics highlight recent inroads made in understanding the genetics of infectious disease susceptibility. The first found an association between risk of meningococcal disease and CFH, a gene previously implicated in age related macular degeneration. The second identified a susceptibility locus for tuberculosis in African samples. Paul de Bakker and Amalio Telenti have a nice News and Views piece about them as well, remarking on this welcome advance not only in understanding infection, but also in using GWAS to gain insight about disease risk in non-Europeans. [JCB]
Update: Dan Frost from the GoldenHelix blog has drawn our attention to a thought-provoking post on the future of GWAS studies. The post suggests that much of the missing heritability in complex disease is hiding in the set of variants that are badly tagged by existing chips, and proposes that GWAS studies in the future may include a sequencing phase to discover new variants in cases, followed by genotyping using custom genotype chips to capture this variation. The question, from my point of view, is how many common SNPs are there that aren’t well tagged by existing chips, and thus how much heritability could be hidden there? This is exactly the sort of question that the 1000 Genomes dataset was designed to answer. [LJ]
 Out in Nature this week is a paper by three Genomes Unzipped authors reporting 71 new genetic associations with inflammatory bowel disease (IBD). This breaks the record for the largest number of associations for any common disease, and includes many new and interesting biological insights that you should all go and read about in the paper itself (pay-to-access I’m afraid) or on the Sanger Institute’s website.
Out in Nature this week is a paper by three Genomes Unzipped authors reporting 71 new genetic associations with inflammatory bowel disease (IBD). This breaks the record for the largest number of associations for any common disease, and includes many new and interesting biological insights that you should all go and read about in the paper itself (pay-to-access I’m afraid) or on the Sanger Institute’s website.

 To celebrate 10 years since the back-to-back publications of complete human genomes in Science and Nature, Science has published series of articles looking back at the last 10 years of genomics, and forward to the future. The article contains short essays from Francis Collins and Craig Venter, the former talking about some of the successes of medical sequencing (including giving a name and photograph to the exome-sequenced IBD patient I
To celebrate 10 years since the back-to-back publications of complete human genomes in Science and Nature, Science has published series of articles looking back at the last 10 years of genomics, and forward to the future. The article contains short essays from Francis Collins and Craig Venter, the former talking about some of the successes of medical sequencing (including giving a name and photograph to the exome-sequenced IBD patient I  Of particular interest is an article by Eliot Marshall on why genomics hasn’t yet had a large effect on medical practice, and what needs to be done to allow the genomic revolution to trickle into medical care. He argues that scientists and doctors need to meet each other half way; scientists need to focus more on showing the direct clinical utility of genomics, whereas doctors need to be more ready to accept new technologies and discoveries, and adapt the way they practice medicine to make full use of them. [LJ]
Of particular interest is an article by Eliot Marshall on why genomics hasn’t yet had a large effect on medical practice, and what needs to be done to allow the genomic revolution to trickle into medical care. He argues that scientists and doctors need to meet each other half way; scientists need to focus more on showing the direct clinical utility of genomics, whereas doctors need to be more ready to accept new technologies and discoveries, and adapt the way they practice medicine to make full use of them. [LJ] There is a real “wow” paper out in pre-print at the journal Genetics in Medicine. It is a wonderful example of the application of cutting edge sequencing technology to solve a medical mystery. Even better, the authors also include an auxiliary discussion about the medical and ethical issues surrounding the diagnosis, which raises some interesting issues about the transition from research to clinical sequencing.
There is a real “wow” paper out in pre-print at the journal Genetics in Medicine. It is a wonderful example of the application of cutting edge sequencing technology to solve a medical mystery. Even better, the authors also include an auxiliary discussion about the medical and ethical issues surrounding the diagnosis, which raises some interesting issues about the transition from research to clinical sequencing. ScienceNews reports that researchers from the University of Copenhagen have got permission to sequence the genome of Sitting Bull, the native American war chief that led the battle of Little Bighorn. I don’t know exactly what they intend to learn from the genome scientifically, but it seems like this might serve primarily as a monument to a major figure in native American resistance. So the question I have is this: how can we go from a genome sequence (which is generally just a text file on a computer) to a public rememberance, something akin to the 1989 postage stamp shown to the left? [LJ]
ScienceNews reports that researchers from the University of Copenhagen have got permission to sequence the genome of Sitting Bull, the native American war chief that led the battle of Little Bighorn. I don’t know exactly what they intend to learn from the genome scientifically, but it seems like this might serve primarily as a monument to a major figure in native American resistance. So the question I have is this: how can we go from a genome sequence (which is generally just a text file on a computer) to a public rememberance, something akin to the 1989 postage stamp shown to the left? [LJ] RSS
RSS Twitter
Twitter
Recent Comments