Last week, scientists at the European Molecular Biology Laboratory reported that they had sequenced the genome of the Henrietta Lacks, or “HeLa”, cell line. This report was met with considerable consternation by those who (justifiably, in my opinion) wondered why scientists are still experimenting on a cell line obtained without consent in the 1950s [1]. In response to a bit of a backlash, the researchers removed the HeLa sequence from the public internet, and even the paper itself might disappear from the formal scientific literature.
However, it is unfair to treat the authors of this paper as scapegoats for the systematic failure of scientists to deal with issues surrounding genomic “privacy”. Consider this important piece of information: the genome sequence of the HeLa cell line has been publicly available for years (and remains so).
This fact is a simple consequence of the fact that nearly every large-scale molecular biology techinique relies on DNA sequencing as a readout. Every time anyone does a genome-scale experiment (for example, RNA-seq or ChIP-seq) on HeLa cells and archives the data, they are explicitly making public the genome sequence of HeLa cells, and thus of Henrietta Lacks.
It’s quite trivial to demonstrate this point. HeLa cells are one of the cell lines used in the ENCODE project, an ambitious effort to functionally characterize the entirety of the human genome. As part of this effort, they (among many others) have been subjected to an array of genomic experiments, almost all of which of course generated sequencing data. I downloaded a few sequencing files from these experiments, and combined the HeLa sequence with other publicly available genome sequences from the 1000 Genomes Project (for full details, see below).
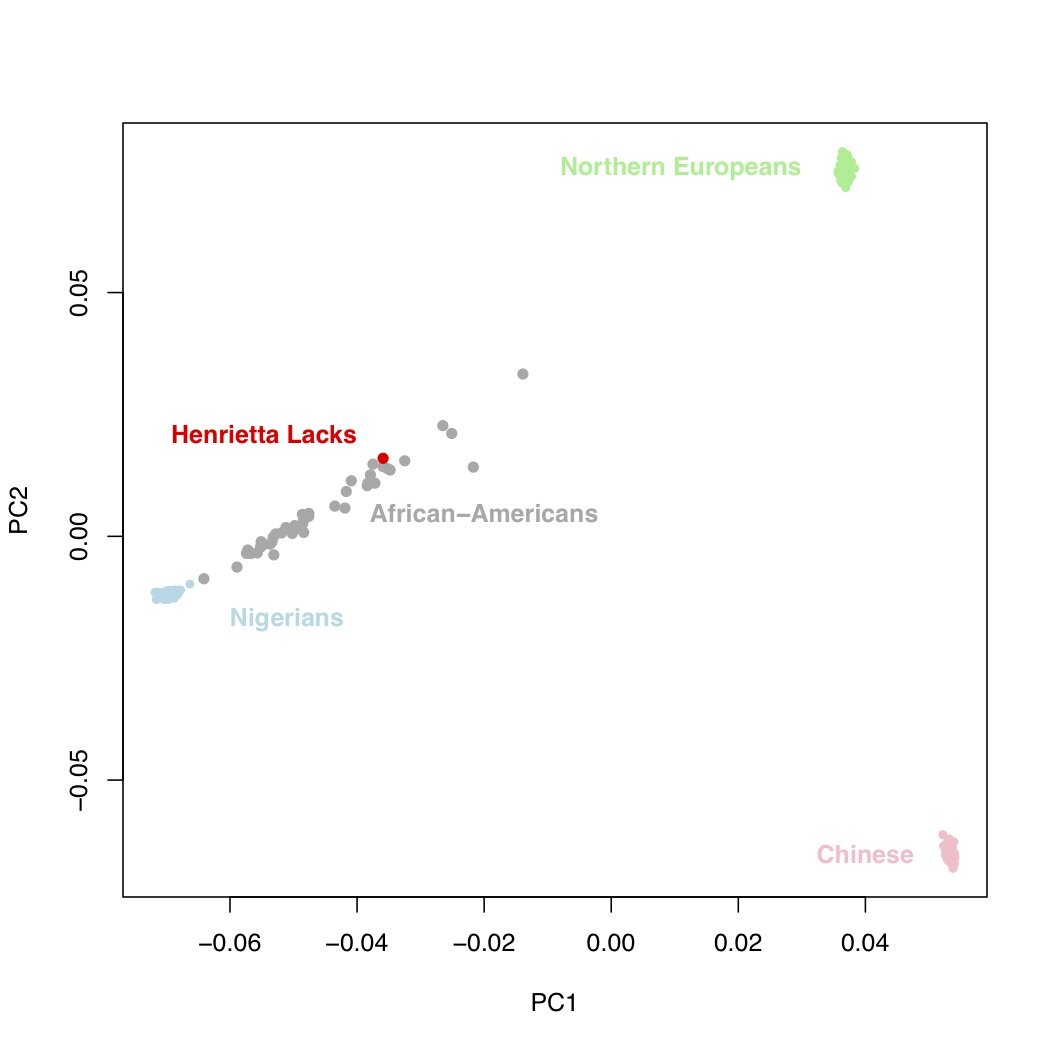
Does the data on HeLa cells contain enough information to say anything about Henrietta Lacks? Plotted above is the output of principal components analysis computed on genetic data from Nigerians, northern Europeans, Chinese, African-Americans, and HeLa. Each point represents an individual, and individuals that fall closer together are more similar genetically. Based on this plot, we can see that the HeLa cells are quite clearly from an African-American woman (or at least someone who is admixed between European and African populations).
These ancestry results are just a proof-of-principle, but any genetic analysis of disease risk or other phenotypic trait is of course just as trivial. This is not true just for HeLa cells, but across the board–the genomes of the donors of every cell line studied by ENCODE are publicly available, and can be analysed for ancestry or disease risk. Though the identities of the donors is not known in most cases besides HeLa, using techniques like those used by Gymrek et al., it may be possible to link cell lines to last names, and thus genetic information to individual people.
[1] See the spectacular The Immortal Life of Henrietta Lacks for the complete back story on this cell line.
—–
Technical details:
I downloaded sequencing data from the following GEO accessions: SRR227441, SRR227442, SRR227445, SRR227446, SRR227472, SRR227473, SRR227505, SRR227506, SRR227556, SRR227557, SRR350914, SRR350915 SRR568260, SRR568261, SRR577378, SRR577379, SRR577392, SRR577393, SRR577429, SRR577430
These are all ChIP-seq experiments on HeLa cells from the ENCODE project.
I then mapped reads to human hg19 using bwa. To compare to the 1000 Genomes data, I used the genotypes from the Illumina OMNI array. To merge HeLa into these data, I randomly sampled a single sequencing read covering each site on the array (at all sites that had at least a single read covering it) and reported the sequence from that read as the HeLa “genotype” (calling heterozygotes is moderately annoying, so I didn’t try it). In total I genotyped 2095422/2177885 (96%) of sites successfully with this approach. I then ran PCA (using smartpca) on the genotypes from the YRI, ASW, CEU, CHB, and HeLa samples.








 RSS
RSS Twitter
Twitter
I wonder why scientists are still experimenting on a cell line that is totally screwed up and keep using it as a model for the normal processes of life.
You make good a good points. The family has asked, per NPR, that the sequence be taken down while they try to understand all of this, which I don’t think is an unreasonable request.
Maybe it is time to let Henrietta rest in peace.
I agree 100% that the authors of this paper should not be treated as if this is an issue that they alone created or were solely involved with. As I said in the OpEd: “The publication of the HeLa genome without consent isn’t an example of a few researchers making a mistake.” This is why I didn’t mention them by name in the article — I did obviously have to cite where the research was done, but I did not want to sidetrack the issue by having the public focus specifically on those individual researchers.
There are, as you point out, several other databases that include HeLa genetic information to varying degrees. But this is the first full HeLa genome publication, it’s also the first of this genetic information published in a format that can be so easily translated into personal information about Henrietta (by scientists and non-scientists alike) with a few mouse clicks online. Since it was the first full genome publication, it’s also the first time any such data got press attention that brought it to the attention of anyone outside the circles of scientists who know and/or use those databases. As Jonathan Eisen has pointed out on his blog and Twitter, the question of what should happen with the HeLa data in these other databases is still an open one. I contacted the director of ENCODE to ask about this and he did not respond to my request for an interview.
Due to newspaper space limitations (alas) I couldn’t go into much detail about this in my OpEd, but I did mention that these other databases contain this genetic information, also without the Lacks family’s consent (in earlier drafts of the story I had a more detailed section about these databases, but unfortunately that was cut for space). Thanks for going into more detail about the data here — this is useful and important information for people to know.
@Rebecca Skloot,
Thanks. I had started to sense a bit of animosity towards the authors of this paper from some comments, but not yours.
There is also this RNA-seq study on HeLaS3 from 2008: http://www.biotechniques.com/multimedia/archive/00001/BTN_A_000112900_O_1994a.pdf
You might also want to point towards SNP chips like this one for example: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM723055
It might not be a “full genome”, but there isn’t really a qualitative difference between genotyping by DNA-Seq and genotyping by array when one is looking for the presence of known disease alleles.
The genome (i.e. the set of SNP genotypes) of a HeLa cell line was out there even before ENCODE.
Excellent post!
For me the issue was the bad press release of EMBL that stated that we can’t learn anything from HeLa-Seq. Your post adds more evidence that this is not true, and even Chip-Seq data is enough to expose the germline variations.
The main point is that genetic privacy is not binary, but a continumm. Having the cell line all over the world diminishes Lack’s genetic privacy, ENCODE FASTA-files furthers reduces Lacks’s privacy, and EMBL VCF almost compromised it totally.
@Yaniv Erlich,
Thanks for the comment. I agree that the press release from EMBL was unfortunate. I’m not so sure about the “continuum” view of genetic privacy, but will have to think about it. For example, imagine a world where every home had a genome sequencer–in that world, having the cell line is essentially equivalent to having the genome sequence.
In this specific case, the genome sequence = ENCODE fastq files + a few hours of time from a computational geneticist. The latter might not be *easy* to come by, but for anyone with a bit of motivation (and maybe a few dollars to spend) it’s within the realm of possibility. And of course now that I’ve put in the time, it would be simple to upload a VCF and move us all the way to the other end of the spectrum.
I guess the counterargument is analogy to other sorts of personal information (e.g. important passwords, social security numbers, etc.) that are somewhat secure because they’re a pain to get without specific expertise?
I think that both Yaniv and Joe are agreeing on a basic point: genetic “privacy” is a function of how many people can see your DNA sequence with a given amount of effort (time/money, where money can presumably buy expertise).
At the moment, putting a person’s DNA sequence on the internet substantially reduces their privacy, especially in a world where few people have home sequencers, access to samples, or the analytical tools to handle the data. Converting the raw data into a “clean” form further reduces privacy by making the information accessible to non-experts. Joe’s hypothetical future would change the equation by democratizing the technology: if anyone can get clean DNA sequences quickly and cheaply, then anyone can breach your genetic privacy if they get ahold of a tissue sample.
From this standpoint, the successive releases of Henrietta’s genetic information (cell line, ENCODE, EMBL) can be viewed as increasingly severe breaches of her privacy in the context of current technology — I think this is what Yaniv meant. As the technology and tools advance, the steps in the middle won’t matter as much, and we will be left with the fundamental problem that a person’s identified biological material was disseminated around the world without her consent.
@Bryan,
Thanks, those are good points. Maybe an operational definition of a “privacy continuum” is: what’s the price (time/money) that a random person would have to put in to find out a target’s (e.g.) APOE status? Targeting me costs ~$0, so I have no privacy. Targeting the donor of an identified ENCODE cell line costs whatever I charge for a few hours of work, so there’s some (unfortunately for my bank account, small) level of privacy.
Releasing any amount of genetic data into the public domain is a calculated risk. As you all of you know releasing genotype data at common SNPs is reasonably predictive at rare SNPs. Even releasing a few markers could carry risks that we don’t yet predict. However, few phenotypes are well predicted by genetics, and risks could be predicted by family history. So there’s a good reason not to be too concerned in the long run, putting aside the specifics of the situation here (which are obviously not good). It would be interesting to know if people hide family history of disease from their doctors, or are unwilling to discuss family diseases in public or on the internet. My sense is the answer may be yes to both questions, but it would be interesting to know whether folks adjust their concern by genetic risk info and family history info in the same way. I’m guessing not, but I would be interested in seeing a discussion of public understanding of risk and relatedness vs genetics. Perhaps this is out there somewhere?
I’m also pretty sure that very few of us have the ability to assess the ability of our relations being sequenced (genotyped). I’ve struggled with this calculation quite a bit for complex family relationships, and I’m not too bad at these things. Putting aside the specifics of the case here, I’m also not sure what degree of relatedness to a dead relative would give me a reasonably claim to have a say in “consent” issues. That calculation is not simple and depends on how I weight different info in a non-trivial ways, so I understand the significant grey area here.
Graham, it’s also the case that there are highly heritable diseases which may be ‘unrevealed’ in family members. A good example is bowel cancer – this can be highly heritable, but there may not be any known family history simply because people died of something else before the condition was manifested in them – maybe developing but symptomless, or maybe not yet developing. People may have such a highly heritable condition without knowing.
@Graham
It would be interesting to know if people hide family history of disease from their doctors, or are unwilling to discuss family diseases in public or on the internet.
This is a good question. Molecular genetic information is not fundamentally different than guessing risk from known cases in pedigrees, though it may feel that way.
I’m also not sure what degree of relatedness to a dead relative would give me a reasonable claim to have a say in “consent” issues.
Agreed. I tried to avoid editorializing too much in the post, but I certainly did not give my extended family veto power over my decision to take part in the personal genome project.
Moving somewhat off topic.
I wonder if there is a way to use people’s intuition about family history to talk more clearly about disease predictions from genomics. That would allow one to move from predictions of disease risk to degree of relatedness of an individual with a disease. E.g. most predictions from knowing your genome are currently worse than the info you learn from knowing the a 2nd cousin has a disease. But some better understood diseases would be like knowing a grandparent has a disease. Such a framework might be more intuitive than straight probabilities, and also (to me at least) it seems to somewhat by-pass the trap of genetic determinism. Or it may just muddy people’s understanding further, and be totally unhelpful.
It might also offer a way to better assay an individual’s understanding and level of consent to having there genome sequenced in the face of different risks. This could be implemented by asking questions about what about family history of disease you would would be comfortable talk to a doctor about, a researcher, a member of the public, etc. However, I’m sure that folks have given this a lot more thought than me, and likely see the flaws in this approach.
“It would be interesting to know if people hide family history of disease from their doctors, or are unwilling to discuss family diseases in public or on the internet.”
At the very least, there is a vocal and identifiable minority for whom this is not not a constraint:
http://www.forbes.com/sites/bruceupbin/2013/03/01/building-a-self-learning-healthcare-system-paul-wicks-of-patientslikeme/
@Graham,
I like that idea, and I actually have no idea about the correspondence between those two things for most diseases. Kind of an interesting thought.
Like is finding out you’re APOE4/APOE4 the equivalent change in risk as finding out a grandparent has Alzheimers? Or a sibling? I have no real intuition about these things, would be a useful exercise to go through the numbers.
If a consensus is forming that consent should be needed from the family of Henrietta Lacks from the genome of her immortal cell line to be published, I think that puts us in worse ethical dilemma than we were already in.
The argument, as I understand it, is that because the genome might reveal (probabilistic) genetic information about the family, they have a right to veto the publication. But today, I can have my own genome sequenced and published without the consent of my family members – only my consent would be needed.
If the families of living people who publish their own genetic information have no rights over the data, why should the families of dead people?
@Richard Smith,
Thanks for the comment. I think maybe the consensus is forming that, since no consent was given by Henrietta Lacks to make a cell line, experiment on it, or sequence its genome, her family somehow has rights (not legal, but maybe moral) to be involved in deciding how it’s used.
I’m somewhat sympathetic to this point of view since this is a bit of a special case–a scientifically important cell line was created in a situation that now would be considered unethical–but I certainly hope it will not be a model for how we deal with these issues in the future.
I’m enjoying the debate about family perceptions about risk.
One of the quotes from Graham: “I wonder if there is a way to use people’s intuition about family history to talk more clearly about disease predictions from genomics”
There is a WEALTH of information published on Lay Beliefs about Genetics and the ‘intuitions’ that people have about how and why diseases are inherited. Most commonly people create their own theories that mean something in the context of their family – ‘my aunt had red hair and was feisty and she had breast cancer, but I have blond hair and am quiet, therefore I’m not at risk from breast cancer’. Or ‘I’m just like my Dad, he has bowel cancer, therefore I’m going to get it too’. These lay beliefs are well documented and any genetic counsellor or clinical geneticist will be able to report numerous families they’ve seen when they’ve spent literally 30 mins describing dominant inheritance in detail and at the end the patient has said ‘yeah, I know you are telling me my risk is 50/50 but I feel and know it is higher because of X,Y,Z’. Lay beliefs are very strong and they don’t necessarily have any logic or scientific basis to them and yet they can be really influential with regards to decision making about testing or screening – or as with Henrietta Lack’s family – their lay beliefs are guiding their need to be involved.
That unfortunate press release indeed contained a huge blunder, as Yaniv and others pointed out. All I can say is that that part of the news release (the Q&A section) initially went out without participation of the PIs. We’re revising procedures at EMBL to avoid such incidents in the future.
Thanks to Joe and other posters on this page for the thoughtful posts.
@Wolfgang Huber,
Thanks for the comment. Yours is not the first unfortunate press release ever made (nor will it be the last); I hope you aren’t getting too much flak for this.
@Anna and @Joe. Yes perhaps people’s understanding of familial disease risk is poor enough that converting calculations from genomic risks back into statements about family history may be totally unhelpful. Even reasonably smart undergrad biology students in my class sometimes say things about the genetics of transmission that make me worry that they’d not handle genetic or family history info well.
DNA sequencing will be commonplace and we need to determine WHO gets to do WHAT with each of our genome sequences. Miinome is claiming DNA sequence is personal property and they want to serve as a broker to bring value to the owners (members) by marketing DERIVED data (not raw sequence)to customers (eg. the rest of the world). Contrast that with 23&Me’s new API that shares ACTUAL DNA SEQUENCE, base by base..which do you prefer? I see this headed to the courts for sure!
not sure we will ever be able to get rid of HeLa cells….its thought that HeLas, due to their aggressive growth, etc, contaminate many established cell lines (e.g., WISH)
http://link.springer.com/article/10.1007%2FBF02616110?LI=true
http://www.archivesofpathology.org/doi/full/10.1043/1543-2165-133.9.1463
http://www.sivb.org/publicPolicy_Eradication.pdf
I agree and would also like to point out that the HeLa cell line can be obtained and sequenced by anybody! Privacy was violated when the original cell lines were established and distributed.
What about the reverse situation? What if the rest of the immediate family wants someone to provide their genome sequence, yet they refuse? Should a genome sequence be expected from a parent, like child support?
For instance, if a mother has had early, aggressive, breast cancer. The children all found that they were not carriers of known causative mutations, yet the grandfather was a carrier. All of the children would like to know if their mother had inherited this gene. Even still, she refuses to be tested. When she dies, they want to have her genome sequenced. Is this ethical? When people have a virus, and may have passed it to other people, they are expected to tell them. Why not with genetic diseases?
This is such a fascinating story. One issue that Rebecca Skloot does not address is that she herself revealed the names and family associations and the origin of the HeLa cells. Certainly these were taken without consent at a time where consent was not considered and the repercussions in terms of genetic legacy or effect on our understanding of biology were not dreamt of. But the identity of the unknowing donor seems to have been a well-kept secret and unearthing the identity of Ms. Lacks took a serious investigative effort. Rebecca Skloot did this at a time when the adverse effects of identifying the family were much clearer. I understand the desire to correct an old wrong and the inquisitive nature of journalism, but I also wonder if Ms. Skloot has not now become complicit in an ongoing tragedy. I’m quite sure the Ms. Skloot had the best of intentions and meant no harm, but her work now exposes generations of Lacks family members to both the shadow of tragedy and the very real issue associated with identified genomic sequence.
I have used cancer cells derived from Ms Lacks. I cannot see any problem, those cells are in effect immortal as they keep dividing provided the cells have sufficient nutrients. That is what makes those cells so important, their immortality whereas the rest of Ms Lacks cells have died with her. I am sure if you collected the cells in pus you could obtain all the genetic information from their nuclei. Is that wrong?
Henrietta Lacks was a one of a kind. Using her perfect cells are not like raiding her underwear drawer. She has saved countless lives because of the research that has led to cures. Let get some perspective here!
It is very short-sighted to be worrying about a long dead woman’s *privacy*. Going forward, consent should be given but when push comes to shove and there is some genetic fluke like Ms. Lacks who could led to more cures, involuntarily acquiring cells through court action, is not out of the question.