Out in the PNAS Early Edition is a letter to the editor from four Genomes Unzipped authors (Luke, Joe, Daniel and Jeff). We report that we found a statistical error that drove the seemly highly significant association between polymorphisms in the OXTR gene and prosocial behaviour. The original study involved a sample of 23 people, each of whom had their prosociality rated 116 times (giving a total of 2668 observations), but the authors inadvertantly used a method that implicitly assumed there were actually 2668 different individuals in the study.
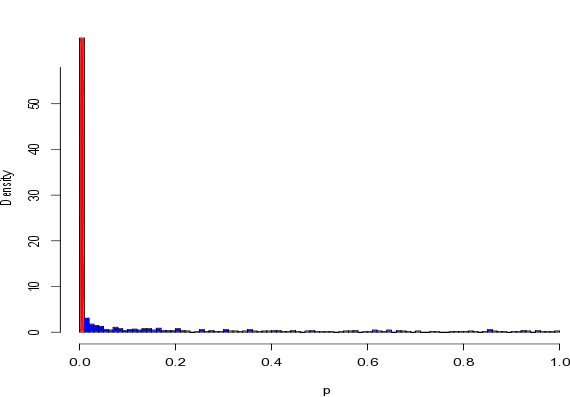
The authors kindly provided us with the raw data, and we ran what are called “null simulations” on their dataset to check to see whether their method could generate false positives. This involved randomly swapping around the genotypes of the 23 individuals, and then analysing these randomised datasets using the same statistical method as the paper. These “null datasets” are random, and have no real association between prosociality and OXTR genotype, so if the author’s method was working properly it would almost never find an association in these datasets. The plot below shows the distribution of the “p-value” from the author’s method in the null datasets – if everything was working properly all of the bars would be the same size:
The very large red bar represents false positives, and you can see that the approach the authors used generates low p-values (and thus seeminly very significant associations) even when there are no associations to be found.
We previously wrote about this study on this blog, and noted the problem with these sort of candidate gene studies, where a very small sample size is used to study a single gene. The power is so low that, even if there was something to be found these studies would not be able to detect it, and as result the p-values they generate are more or less random either way. The same argument also explains why these sort of studies are enriched for statistical mistakes – there is no power to find a true association, so a low p-value is more likely to be caused by an accidental slip-up with the statistics than by a real association. Without statistical power you are fishing without bait, and a big tug on the line is more likely to be a shopping trolly than a fish.








 RSS
RSS Twitter
Twitter
Well spotted. I thought something like this must have happened when I read the paper, but I wasn’t sure of how best to prove that statistically so I never followed it up. Thanks for doing that! An important lesson for behavioural geneticists.