 Konrad Karczewski and colleagues from Stanford have put together a very handy set of online tools for analysing personal genomic data. The tools work within your browser (Chrome and FireFox only, so the ~18% of you who continue to use Internet Explorer now have yet another incentive to change), meaning your genetic data never actually leaves your computer. They currently work with raw, unzipped data from 23andMe and Lumigenix. The tools were developed initially for use in Stanford’s pioneering Genomics and Personalized Medicine elective course for graduate and medical students, in which students had the opportunity to explore their own 23andMe or Lumigenix data. Karczewski has some background over at his personal blog.
Konrad Karczewski and colleagues from Stanford have put together a very handy set of online tools for analysing personal genomic data. The tools work within your browser (Chrome and FireFox only, so the ~18% of you who continue to use Internet Explorer now have yet another incentive to change), meaning your genetic data never actually leaves your computer. They currently work with raw, unzipped data from 23andMe and Lumigenix. The tools were developed initially for use in Stanford’s pioneering Genomics and Personalized Medicine elective course for graduate and medical students, in which students had the opportunity to explore their own 23andMe or Lumigenix data. Karczewski has some background over at his personal blog.
Once you’ve pointed Interpretome to your raw data file (top right-hand corner) and assigned your ancestry you can start playing with the tools – for instance, you can calculate your type 2 diabetes risk or warfarin dose, or estimate the fraction of your genome inherited from Neandertal (see image above for my result). A caveat: I’m writing this post without carefully checking the output from any of these analyses, so as always in personal genomics, interpret your results with caution.
I suspect one of the more popular suites of tools will be the PCA package, which allow you to place your genetic data in the context of worldwide patterns of genetic variation. Here the authors have pre-calculated the crucial information (the PCA loadings) for each SNP in the 23andMe data-set, allowing them to very quickly calculate your position in a worldwide genetic map containing thousands of individuals. Here’s my 23andMe v3 data (black square) projected onto a genetic map of Europe created with POPRES samples. The picture isn’t quite as pretty as the one in the 2008 Nature paper using the same cohort – the Interpretome team haven’t applied the same extensive filters to remove extraneous features from the data have had to work with the smaller number of SNPs that overlap with the 23andMe v3 chip, and you need to plot PC1 vs PC4 before you start seeing something that resembles a map of Europe – but it’s enough to give you a sense as to where you fit. I was unsurprised to find myself sitting smack in the middle of the British cluster:
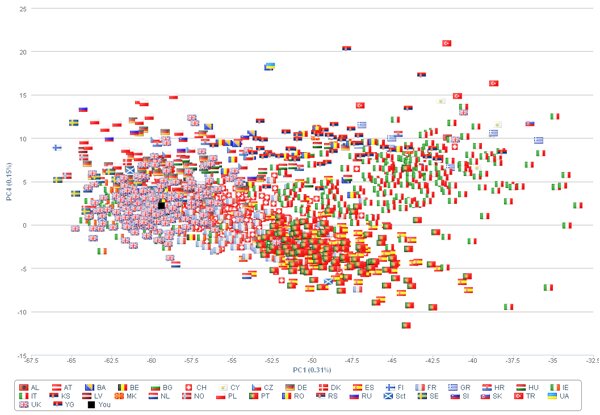
Anyway, go and check it out, and send it to your friends. We’re delighted to see such a handy package released free to the public – kudos to everyone involved in putting the website together. We’ll likely be posting a more thorough review of the site once we’ve had time to test the tools out on a range of Genomes Unzipped data-sets.








 RSS
RSS Twitter
Twitter
The “explore” tab is an absolute mess and crashes for me using Chrome. I also found it impossible to run the explore analyses on my uploaded data; it keeps prompting for upload (in 2 different places on page?) then complaining that there are “too many SNPs” or worse, doing nothing.
The other tabs all work fine and provide interesting visualisations/data.
The chromosome painting uses acronyms I can’t decipher (what ancestry is GIH? TSI?) and when I tried sending an email to the feedback address to ask about it, the email bounced. Neat tool otherwise.
Q: what ancestry is GIH? TSI?
A: http://www.sanger.ac.uk/resources/downloads/human/hapmap3.html
Thanks!
As an East Asian, I notice that while they allow you to select your ethnicity in the upper right, this setting doesn’t seem to be used in any of the risk analyses, nor is applicable ethnicity ever noted in the SNP annotations. Granted, I only looked briefly so it’s possible I am not seeing something. The only mention of this issue I saw was in the text, where it says something along the lines of “not every annotation is applicable to every population”.
I wonder what the FDA would think of this tool? ;)
@Neil: The genotype loading happens in the top-right corner. Any exercises you load would be in the “Explore” tab. If you loaded your genotype into there, it will load, as it takes a similar format, but it will crash since you are trying to compare yourself to yourself. Look at the “Example Annotation File” to see what kind of annotations you can load. I suggest the other exercises as a place to start.
@Nox: Apologies for the bounce. Fixed that typo. Kasper has got it right.
@shwu: Overall, we use the ethnicity for the imputation framework and it also sets the prior and which SNPs are chosen for the diabetes risk calculation. You are right, for the exploration analyses (which follow the lectures given in the course), the ethnicity is not necessarily given. Thanks for the suggestion!
Konrad,
I used you Ancestry Painting feature. I’m 100% Irish.
Using HapMap 2, I was 100% Red=CEU
Using HapMap3, and setting the threshold to 40 SNp’s, I am a mosaic??
HapMap3-Irish-pconroy
The problem with their approach is that you have include 2 recently Admixed populations – African-Americans and Mexican-Americans – and you have no West Asian reference population??
So the African-Americans hits I have are probably just Northern European – as they don’t have a Northern European reference population, for instance!
While the Mexican-American hits I’m getting probably represent Western Europe/Atlantic Europe, and not necessarily Maya or Pima like sequences.
Where the Gujarati probably represent a West Asian component best (ANI)??
What do you think?
@pconroy: You are right, the Hapmap3 ancestry painting still needs a bit more development. We put it in for experimental purposes, but as you said, since there are 2 admixed populations, you are likely to see patterns of admixture from those populations (since, of the options given there, Europeans will resemble Italians most, followed by Mexican, Indian, or African-American). We are working on some more distinct reference populations and will update as they become available.
Every weekend i used to pay a visit this web page, as i
wish for enjoyment, as this this site conations really nice funny information too.
I enjoyed exploring a new approach to DNA, but I didn’t know what the initials of many of the answers meant. I showed no Neanderthal. On 23andMe I am 2.7%. Was I supposed to download additional information or did I not do something?