UPDATE 3/17/12: A more extensive analysis of the paper discussed in this post is here. Several groups have concluded that at least 90% of the sites identified are technical artifacts
The “central dogma” of molecular biology holds that the information present in DNA is transferred to RNA and then to protein. In a paper published online at Science yesterday, Li and colleagues report a potentially extraordinary observation: they show evidence that, within any given individual, there are tens of thousands of places where transcribed RNA does not match the template DNA from which it is derived [1]. This phenomenon, called RNA editing, is generally thought to be limited (in humans) to conversions of the base adenosine to the base inosine (which is read as guanine by DNA sequencers), and occasionally from cytosine to uracil. In contrast, these authors report that any type of base can be converted to any other type of base.
If these observations are correct, they represent a fundamental change in how we view the process of gene regulation. However, in this post I am going to point out a couple of technical issues that, if not properly taken into account, have the potential to cause a large number of false positives in this type of data. The main point can be summarized like this: RNA editing involves the production of two different RNA and/or protein sequences from a single DNA sequence. To infer RNA editing from the presence of two different RNA and/or protein sequences, then, one must be very sure that they derive from the same DNA sequence, rather than from two different copies of the DNA (due to, for example, paralogs or copy number variants). Although this issue has the potential to be a large source of false positives in a study like this, I will discuss an additional technical problem that could also result in false positives.
How to identify sites of RNA editing?
The experimental approach of the authors is easy to explain: they sequenced the mRNA expressed by an individual (or rather, cDNA from a cell line derived from the individual), obtained DNA sequences from the same individual, and compared the two. Any difference between the RNA and DNA should be due to RNA editing.
The complication comes from the fact that, with current technology, it’s impossible to sequence an entire mRNA or genome in a single pass. Instead, what the authors have are short reads (of 50 bases) from the mRNA of the individual, and reads of various length from the DNA of the individual (from the 1000 Genomes Project). What they have to do, then, is match these sequencing reads to the genome (or transcriptome) to see where they came from.
It is this step that can cause problems in an analysis like this. The authors limit themselves to considering positions in the genome where reads map uniquely; that is, reads that match a single position in the genome better than any other position (Actually, Li et al. (2011) map reads to the transcriptome—the regions of the genome annotated as expressed. This adds additional complications in this step, but for most of what follows these complications are not relevant). However, having a read map uniquely is far from a guarantee that it’s in the right place—choosing the best spot involves a number of assumptions, including how you weight insertions and deletions and possible sequencing errors. How could this influence the results in this paper? Let’s take a couple examples:
Paralogs can confuse read mapping
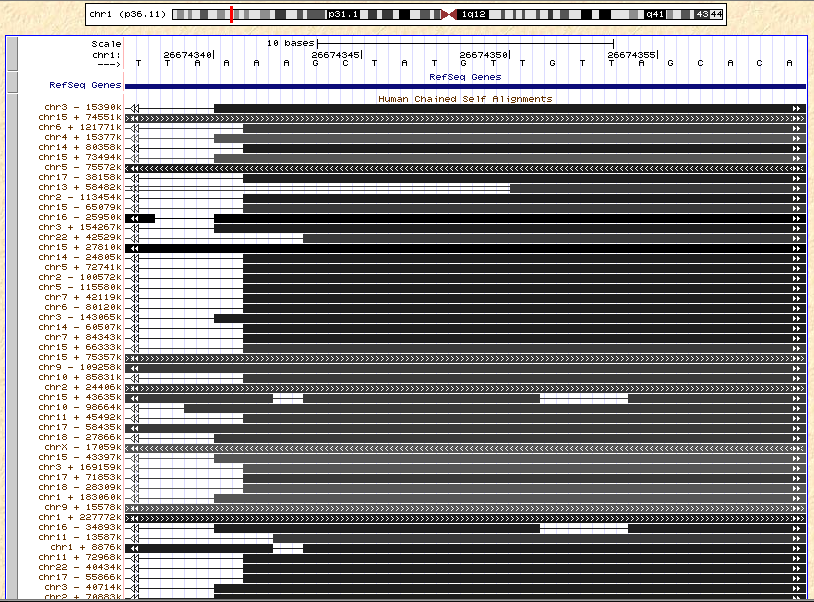
One example of a potential RNA editing event is in the gene HMGN2 (from their Table 1). Below, I’m showing a screen shot from the UCSC Genome Browser in the region of the putative site (you may have to open the image in a new window to get it to a readable size). What I’m showing is the DNA sequence, the position of the RNA (the putative editing site is in position 26,674,349, where the authors claim the G is sometimes converted to a T), and, crucially, a track showing other places in the genome that are similar to this region. By my count, there are 62 other regions of the genome with high percent identity to this region (in the screen shot below, some of these are cut off). To be sure that the T allele at this position is not from any of these other 62 regions, it would be important to show evidence that 1) these 62 regions are not expressed (inspecting some of them shows that some are indeed genes), or 2) if some of them are expressed, that the sequence in the corresponding position in that region is not a T.
These sorts of mapping issues are well appreciated in the literature on SNP calling from sequencing data (which is another situation in which one is looking for a difference between a sequencing read and the genome); a naïve SNP caller that just looks for differences between aligned reads and a genome will output tens (or probably hundreds) of thousands of false positive SNPs which must be filtered out by various criteria [2]. For a list of criteria that are used for this filtering step, I suggest the paper published by the 1000 Genomes Project [2] (see the Supplementary Information of that paper). As far as I can tell, Li et al. (2011) did not remove false positive calls by any of the criteria generally used in this closely related problem.
Does this matter? I examined some of the reads supporting “editing” at the site mentioned above from the data in Li et al. (2011) (GSE25840). For example, take a read with the sequence TTTTTTTTTTTAAAAGCTATTTTGTTAGCACACAGAACACTTCATTGTTG. In the alignment file I downloaded, this read is mapped over the edited position with extremely high confidence (an alignment quality score of 255). However, you can take that sequence and match it to the human genome using BLAT (just copy and paste into the box after opening that link). There are at least 20 or 30 places in the genome that match that read nearly as well as the “best” match covering the edited site. It’s unclear why this is labeled as a confident alignment, but clearly it should not be. Any errors like this in mapping to paralogous regions can generate false signals of RNA editing. [NOTE ADDED 5/26/2011 by JP: See comment regarding read mapping here. The read mapping software used by the authors gives a mapping quality score of 255 to all reads which map uniquely to the genome, regardless of how unique the read actually is.] For this particular example, it seems possible that in this individual either 1) the relevant genomic base is actually a T at one of the 20-30 regions which closely match the sequencing read (due to either an unannotated SNP or error in the reference genome), or 2) there is an additional copy of the region, absent from the reference human genome, which carries a T at the relevant position [3].
Strikingly, many of the examples the authors give in their Table 1 fall in regions with high percent identity to other parts of the genome, where these sorts of false positive calls are known to be rampant in SNP calling. An interesting example in which this is not the case is below.
Splice junctions cause read mapping problems
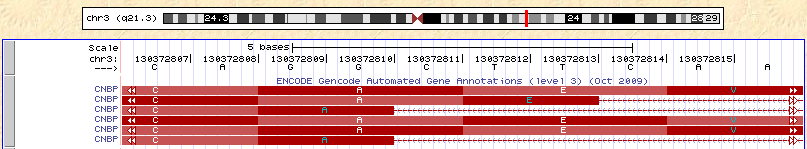
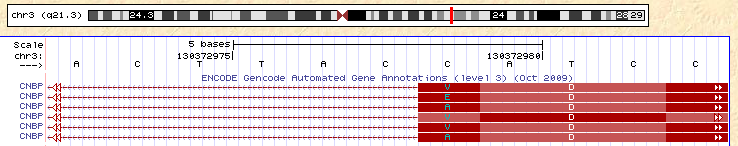
The authors point out that a number of the places where RNA and DNA don’t match fall near splice sites, and they speculate that RNA splicing induces editing. Let’s look at one such example, in the gene CNBP (from their Table 1). This position does not appear to have high percent identify to anywhere else in the genome. Below, I’m showing a couple of screen shots from the UCSC Genome Browser surrounding this splice site (the putative editing site is a T->A change at position 130,372,812). You can see that there are a number of different annotated splice sites at the 3′ end of the intron (the top picture), which are all spliced to a single 5′ splice site (the bottom picture). Read off the sequence of the shortest of the isoforms (the 3rd and 6th from the top)–the sequence of the gene goes CAGG|CATC, where the bar represents the position of the intron. Now consider reading through that splice site (for example, on the top isoform)–the sequence is CAGGCTTC. Imagine you had a read with the sequence CAGGcatc (a perfect match to the short isoform) and tried to map it. The mapper has a choice: either use a gap in the alignment to map across the intron without any mismatches in the read (this is the correct alignment), or map without a gap, but with a single T->A mismatch (this is an incorrect alignment). Many mappers will choose the latter, and this misalignment will cause a false signal of RNA editing.
Again, I found one of the reads in the data from Li et al. (2011) supporting the “edited” site discussed above. This read has the sequence TGCAGTCCTTGGCAATGTGGCCACCTCTACCGCAGTTATAGCAGGCATCC (again, you can map this read to the genome using BLAT), and again has an extremely high mapping quality of 255. I’ve italicized the sequence around the putative editing site. This read is an exact match to the short isoform (with no mismatches), but is mapped with two mismatches and no gaps to the longer isoform by Li et al. (2011), causing a spurious signal of RNA editing. The evidence for RNA editing from this sequencing read is a consequence of the “preference” of the read mapper for mismatches over gaps in alignment. Because of this effect, I’d be particularly suspicious of putative RNA editing events that fall near splice sites.
Conclusions
In this post, I’ve discussed a couple of technical issues that could lead to false inference of RNA editing from data like that used in Li et al. (2011). First, mismapping of reads in paralogous regions could lead to false signals of RNA editing. These false signals would even be replicated in follow-up experiements like those done by Li et al. (2011), because the two forms of RNA and protein are indeed present in the cell. However, the two forms of RNA and protein do not come from the same DNA sequence, and thus are not evidence of RNA editing. Second, mapping biases around splice sites (and other sorts of insertions/deletions in the genome) will cause mismapping and false inference of RNA editing.
So are all the results in Li et al. (2011) due to effects like those above (apart from the A->G editing, which is known to be common)? I can think of a number of analyses that might shed some light on this. First, do most putative RNA editing sites fall into regions with high homology to elsewhere in the genome or near splice sites, like I described for HMGN2 and CNBP above? Looking at Table 1 in Li et al. (2011), I see only one potential example that doesn’t, and it’s an A->G site in AZIN1, but it would be worth examing this on a global scale. Second, what about the biases described by people working in SNP calling—do putative edited nucleotides show a strand bias (do only sequencing reads mapping to one strand or the other show the editing event?) or position bias (do the putative edited nucleotides tend to fall near the start or end of the sequencing reads)? If biases like these are absent, perhaps these technical issues do not have a strong effect.
Finally, I should reassert that RNA editing is indeed an interesting and potentially important phenomenon in humans; that said, skepticism of some of the claims in this paper seems warranted.
—-
[1] Li et al. (2011) Widespread RNA and DNA Sequence Differences in the Human Transcriptome. Science. doi: 10.1126/science.1207018
[2] The 1000 Genomes Project Consortium (2010) A map of human genome variation from population-scale sequencing. Nature. doi:10.1038/nature09534.
[3] Shouldn’t the genomic DNA sequences from this individual control for this? Not necessarily. The sequencing reads generated from genomic DNA by the 1000 Genomes Project have different lengths than those generated from the RNA by Li et al. (2011). Even subtle differences in the mapping properties between the RNA reads and DNA reads will lead to a subset of sites in the genome where RNA reads map well and DNA reads do not (or vice versa), particularly in multi-copy regions.








 RSS
RSS Twitter
Twitter
This is a great overview. I am actually using SNP calling to identify RNA editing in a sequencing project in my lab right now. I’ve been using the SNP calling methods listed in the 1000 Genomes paper and focusing on the A->G and C->T edits for exactly the reasons you detail in your post. It seemed obvious to me, but I guess it wasn’t to the reviewers?
Ditto Josh, great! I suppose this will be another Science paper languishing online before (if ever) getting to the print version…
I don’t pretend to understand all the details, but it seems that the simplest explanation is the paralog issue.
Is another possibility that DNA -> RNA is has less fidelity than DNA -> DNA? This would introduce random differences which would be individually rare, but if there were “hotspot” sequences where bases are more often miscopied (for local sequence/structure reason) which could lead to the accumulation apparently edited RNA molecules. Such sequences exist in DNA – I need to find the refs when I get off the train to a stable connection (quick e.g. http://bit.ly/jwGzLk)
Outstanding review and description of the issues and artifacts related to this analysis.
Great review and analysis of the possible shortcomings in the nucleic acid analysis. But then how do you explain the fact that the peptides isolated in mass spec match the supposedly edited sequences? I am not a mass spec expert, and I know that mapping peptides brings its own unique set of questions. I’d like to see that angle addressed . . .
Hey Joe H.,
Joe P.’s point is that the supposedly “edited” sequences likely come from other transcribed regions of the genome, e.g. closely related genes – so in many cases you would expect to see a mass spec signal for the “edited” transcript, which is actually just a perfectly normal protein isoform produced by a separate gene.
If it’s an artefact of mapping or genome structure, it should be statistically and globally testable using the data sets provided rather than individual examples. I would be interested to know if the cDNA step can create any such errors in reverse transcription. I recall DNA polymerases slipping in sequence context specific ways, under certain conditions, at frequencies within the expected mutation ranges.
Clive makes a good point – cDNA generation will not have the same fidelity as either genomic DNA replication or RNA transcription – it’s something that could be tested.
Also I note that the authors cannot yet say whether the base changing event happens during or post-transcriptionally so it is maybe a bit early to assume it is due to RNA editing:
“For most of the cases, we do not know yet whether a different base was incorporated into the RNA during transcription or if these events occur post-transcriptionally”
Well I wish the authors well but if there isn’t a lot of artifact in their experiment then I wonder how much artifact is in everyone else’s.
People are going to have to go back and check a lot of Northern blots.
Excellent analysis. However, the authors did perform Sanger sequencing of gDNA and RNA from the same individuals to confirm their predicted editing events. If these events are artifacts due to paralogous transcripts, shouldn’t the signal be picked up by gDNA sequencing as well?
@Y.X
However, the authors did perform Sanger sequencing of gDNA and RNA from the same individuals to confirm their predicted editing events. If these events are artifacts due to paralogous transcripts, shouldn’t the signal be picked up by gDNA sequencing as well?
This is a very important point, thanks for bringing it up. The authors have sequenced gDNA and RNA using the same primers for a subset of their sites. Results from this Sanger sequencing are presented in Figure S3, which I have uploaded here for people who do not have access to it. Readers following this discussion, please do follow along on the figure. I have two points to make:
1. There are six sites presented in this figure. 5 of them are A->G editing (the sequence reads T->C on the opposite strand seen in the figure). A->G editing is common in humans (see, for example, here), and I see no reason to dispute these examples. The last example is a potential G->A editing site in PPWD1. Look carefully at those sequence traces. I see some evidence of an A in the genomic DNA sequence (a slight peak shifted somewhat to the right in the figure). Additionally, G->A sequencing errors are common in Sanger sequencing (see here). To summarize this point, the authors present evidence of only one example of a potentially new type of RNA editing, and this example deserves some scrutiny
2. Sequencing of gDNA and RNA is a valid way to confirm RNA editing, but only for single copy regions of the genome. Imagine there are 50 copies of a sequence in the genome, 49 of which carry a G at a site, and 1 of which carries an A. If you sequence the gDNA, you will see only a tiny, probably negligible signal from the A. Now sequence the RNA. The key point is, what you see depends on which sequences are expressed. Imagine that of those 50 sequences only 2 are expressed, but imagine that 1 of the expressed sequences is the one that carries the A. In the RNA sequence, then, you will see a strong signal from the A allele (50% of the total signal), and you might infer that you have confirmed RNA editing.To summarize this point, the paralog issue raised in the main post can also confound this type of validation exercise.
Really well written critique.
If they see the DPP alleles in a previously seen EST that would shore up evidence for editing at that site. However, it is crucial that the alleles not only map exactly to an EST where the ‘edited’ base pair lies, but that the EST maps back to the gene where the genotype is made unambiguously. Row four has an EST that maps almost just as well to two different paralogs with very similar BLAT scores. The EST for TRIT1 (BF367406) doesn’t even seem to BLAT to that gene. This type of BLAST to EST analysis does not protect against the paralog issues you mentioned. In addition, looking at the at table S3 row 2: DCP1, the edited position: (chr3:53,297,343) falls in an intron where the ESTs do not seem to map to (DA808260 and BX100388). This is probably called from the splice-junction error you were referring to; however the fact that the putative RNA-edited base doesn’t even map to the ESTs is a bit disconcerting. I could be doing something wrong however. Would like to be able to scrape S3 from the PDF and do something more systematic.
Also possibly relevant:
http://www.nature.com/nature/journal/v463/n7280/full/nature08909.html
Review Article Expansion of the eukaryotic proteome by alternative splicing
Sorry if this is a dumb question, but I’d be interested to hear from people with more wet lab experience than me to know just anecdotally how consistent these findings are with their experience in the lab. i.e. how often do you sequence a transcript or do a mass spec and have it not match the genome?
Nice input. I would like to raise a concern however. Been thinking about RNA editing myself, and have desperately been trying to find a reference for the “fact” that A->G edits are common. If no genome wide analysis has ever been attempted prior to now, how can we say that? Might be some solid data supporting this claim (and I would highly appreciate being directed to them), but indicating high proportions of other events does not raise a red flag in my head. All the papers that state that A->G is the major even without referencing it does however. But since I’ve tried looking for editing myself I know that there a lot of things to consider.
My guess is that many sorts of editing events occur, but I would also assume that most of what is being published right now contain a lot of false positives. Baby steps.
@Boel
Genome-wide analysis of RNA editing has been attempted before using EST databases and full mRNA sequences. For example, Levanon et al. (2004) identified about 12,000 A->I sites. Their clever trick for removing false positives was to use the fact that the enzyme that catalyzes A->I editing operates on double-stranded RNA, so they looked only in regions that they predicted to be double stranded in vivo
See also Athanasiadis et al. (2004) and Kim et al. (2004).
@ Joe
Yes, I’ve seen the papers by Levanon et al. This is the typical example of what I brought up. A->G events are assumed to be real and other events are just ignored. And sure, we know the mode of action of the enzyme in this case which makes the search easier. Once we know the modes of action for other editing enzymes the search for other events will be easier. If a thorough analysis was conducted on this EST material looking for other events as well we might get very lucky. The Li et al papers makes some bold claims, and might not have the data to back it up completely. I’m just indicating that (for what I know) the assumption that A->G events are the main editing event might be just an assumption, with no real data backing it up. But please correct me if I’m wrong.
Oh, I see, I misunderstood the question. The question is: a priori, is it plausible that undiscovered forms of RNA editing (besides A->I and C->U) exist? Yes, of course, but not all plausible things are true!
Hi Joe… re the mas spec side.
It is actually possible to get statistically significant peptide mass spectrum matches that have no evidence the actual amino acid substitution.
The authors could have included the mass spectra in the supplement where they “hand sequenced” each matching peptide, showing evidence of the amino acid substitution in the b-ion and y-ion series. From the small example in the manuscript, it looks like a good peptide spectrum match, but which of those b and y ions provide evidence for the editing site?
If they wanted to be even more careful, they could have also argued that there wasn’t another peptide with a fragmentation spectrum (and matching precursor mass) that matched closely as well (quite suspiciously they used very large precursor mass windows on their fragmentation spectrum search). Also, using high resolution high mass accuracy fragmentation spectra would have helped here too by targeting the specific precursor ion in their instrument runs. They could have also used synthetic isotope labeled internal standard peptides to further argue these matching peptides are in their data.
A Mascot peptide database search isn’t enough to argue unequivocally that a single amino acid substation is in the data. It certainly points you in the right direction, but careful analysis of individual spectra is really necessary, especially for an idea as remarkable as this.
Zia,
Thanks for the comments. I have no experience with mass spec, so I have no ability to judge their results in this area. This is really helpful.
From the small example in the manuscript, it looks like a good peptide spectrum match, but which of those b and y ions provide evidence for the editing site?
What’s interesting about this example is that the putative editing site is not actually in the peptide. The “editing” site leads to a loss of a stop codon, and what the authors actually show is a peptide corresponding to translation downstream of this stop codon. Assuming this peptide is real, this is only very indirect evidence of any sort of editing.
Hi Joe,
I confused the peptides in their supplement with the small example. The example in C (from their B-cells) is actually a pretty bad mass spectrum. You only get y-ion series and on top of that only 3 matching singly charged y-ions.This is compared to the example above where you get both b-ion and y-ions which makes the match less ambiguous.
Also isoleucine and luecine have the same mass so leucine and isolucine containing peptides will create the same spectra.
Zia
Excellent analysis. I jumped straight to this blog after reading the Nature editorial.
I personally think the library prep step for the mRNA-Seq (cDNA prep) is a potential source of quite massive error, especially when applied across the whole genome. Even the highest fidelity RT enzymes will incorporate errors every 1 – 5 x 10(4) bases or so in vitro. This is not usually a problem for a typical RT application, such as real-time qRT-PCR, where short fragments are used and amplification errors would presumably be random (ish) and likely ignored due to probe/primer mismatches. However, if we assume a human genome size of 3 x 10(9) bases, then we can see that with the error rates above, there is the potential to generate 60,000 – 300,000 base errors if you compare cDNA to the genomic DNA.
In the article, they claim to have found 10,000 “edits”. Hmmmmmmm
Disclaimer: this is a quick and dirty analysis and my numbers could be off.
@Zia
Any desire to write a post explaining how one goes about interpreting mass spec data? :)
@TheDuke
I agree that RT errors (and sequencing errors, as described by Lior Pachter in the Nature News piece) have the potential to generate false signals of RNA editing. I don’t think this is the major explanation for the authors’ results, though–they are only looking at sites where at least 10% of the reads mismatch the genome. This would imply a 10% error rate, which is likely only in some very pathological sequence contexts (though it will happen at some sites by chance). And these errors, of course, cannot account for the mass spec results in the paper. Based on the examples in this post (and looking more systematically, which I’m thinking about describing in a future post), my feeling is that most of the results are due to mapping errors.
It is indeed a very interesting analysis. We have RNA-seq Illumina reads here at the lab and have hard time removing all false positive due to paralogs, pseudogenes and alignment extension over intronic regions occuring around splice junction sites.
From a purely technical point of view, there is something that seems strange to me. Normally, with such a read length, it is not possible to have a mapping quality of 255. With our own sequences (50bp paired-end reads), the maximum mapping quality never exceeds 60. Even with long 454 reads (more than 500 bp), we never reached the value of 255 for the mapping quality. Actually, the value 255 is used when mapping quality is not available. Quoting from the SAM 1.4 specification format (available on samtools website):
MAPQ: MAPping Quality. It equals −10 log10 Pr{mapping position is wrong}, rounded to the nearest integer. A value 255 indicates that the mapping quality is not available.
I do not know how exactly these sequences were mapped on the human genome, but saying that a read is uniquely mapped when the mapping quality is equal to 255 seems very optimistic to me (if the alignment you downloaded is actually in the SAM format).
David,
Great catch, thanks! I had not noticed this, but the mapping qualities in the .sam files I downloaded are either 0,1,3, or 255. The authors used bowtie for alignment, and a bit of googling reveals that (as of 11/2009, see the comment by one of the authors of bowtie in the linked thread) bowtie gives a mapping quality score of 255 to reads that map, and a mapping quality score of 0 to reads that do not (not sure what 1 and 3 are). So I think that solves the mystery of why the reads in my examples are mapped confidently–the authors are using a mapping program that does not distinguish between confident and unconfident alignments.
Joe,
Thanks for the additionnal info about Bowtie. When we first mapped our RNA-seq reads with TopHat (which makes use of Bowtie), we noticed that the problem of reads overlapping intronic regions is rather important, and induces many mismatches around splice junction sites. Additionnally, since TopHat (and Bowtie) does not handle gapped alignment, anytime a known indel should appear in the alignment, several mismatches appear instead because all bases located after the indel are shifted from their correct positions.
We have switched back to BWA. It handles gapped alignments and assigns a 0 mapping quality value to reads having multiple matches. Even if the problem of reads overlapping intronic region is still present, it is lower compared to the alignment we have with TopHat.
Hope this helps
@David
“We … have hard time removing all false positive due to paralogs, pseudogenes and alignment extension over intronic regions occuring around splice junction sites”
And to think that you could have published in Science…
Fantastic post and the follow-up comments by others are extremely informative. I would love to see a post on the mass spec interpretation.
Thanks for the awesome posts here Joe ( I confess I don’t keep up with all of it). Everything gets pushed into Instapaper and I manage to read some of them.
Controversy about this paper via genomeweb:
http://www.genomeweb.com//node/970035?hq_e=el&hq_m=1019462&hq_l=1&hq_v=4b867261b4
Similar analysis results were published last year suggesting “RNA editing of nuclear transcripts in Arabidopsis thaliana” by Meng et al. and nary an eyebrow was raised. Perhaps violations of the Central Dogma stir less controversy when they occur in flowering plants.
“Perhaps violations of the Central Dogma stir less controversy when they occur in flowering plants.”
And when they’re published in BMC Genomics as opposed to Science…
Great analysis Joe. I hope you’ll submit a comment to Science, this criticism deserves a more official status.
I find it shocking that they spend only half a sentence discussing the possibility of mapping errors. It is (or should be) obvious to everyone working with these types of sequencing data how common they are, and how vulnerable their analysis is to mapping problems.
When we want to confirm rare events (e.g. somatic mutations and non A-to-I editings), we may not trust even the most accurate glocal short-read mappers. We should try a local aligner such as ssaha2/bwasw/cross_match/megablast. Local alignment is conservative, but for rare events, it would be good to stay conservative.
@David
BWA and Bowtie are both belongs to the group of ‘unspliced read aligners’, which align reads to a reference without allowing any large gaps.
For Tophat, reads can be aligned to the entire genome, including intron-spanning reads that require large gaps for proper placement.
ref: http://www.nature.com/nmeth/journal/v8/n6/pdf/nmeth.1613.pdf
@Heng Li
Nucleotide local alignment will provide extremely high false positive hits, due to the small size of dictionary, repeat region etc. That is why I am dubious in confirming the rare results by BLAT alignment.
@Joe
Since Bowtie does not yet report gapped alignments, I do not think the alignment would prefer gap mis-alignment instead of non-gap perfect alignment.
@Yichuan
Yes, it’s true that tophat/bowtie does not allow gaps. My interpretation of the example in the post was that it preferred to map without a gap to an exon, rather than mapping over the splice junction. This is a preference for an ungapped alignment in some sense, if you consider mapping over a splice junction a gapped alignment where the possible gaps are already known, rather than discovered by the aligner (ie., they’re determined by the known introns).
I’m not sure about the inner workings of tophat, but this preference is what would happen if the algorithm first mapped reads to the genome, then took the reads which didn’t map and tried to map them across introns–the read would map with mismatches in the first step (causing a false signal of “editing”) and the algorithm would never get a chance to “see” that there would be a perfect alignment if it looked across the intron.
For what it’s worth, the above speculation about the inner workings of tophat is indeed what I’ve seen anecdotally when hacking together gapped alignments myself (Pickrell et al. 2010). If I first map reads to the genome, then discover splice junctions, then try to map all the reads again over the splice junctions, there are a number of reads which initially mapped to the genome (with mismatches) but match better to the discovered splice junctions. I assume the alignment over the splice junction with no mismatches is the correct alignment, and the mismatched alignment to the genome is an incorrect alignment.
@ Joe. Yup, this is what I’ve seen as well. And it is how TopHat works – reads are first mapped to the genome using Bowtie, this creates “islands” of pileups, and then tophat tries to join them using the initially unmapped reads. Would be a really good idea to re-align the reads placed upon exon-junctions, this is a really common problem in RNA seq overall, and gives less coverage for the actual junctions.
@Yichuan
I have never heard that local alignment produces “extremely high” false positive rates. For RNA-seq, I believe it is quite the contrary. I would suggest you map the RNA-seq reads in the GEO alignment with bwa-sw and then call SNPs. The result IS vastly different. Anyway, a true SNP should be robust to the mapper in use. Why not try a few and see what you can get? A few groups who want to confirm somatic mutations do use multiple mappers (e.g. maq+ssaha2; bwa+mosaik).
Thanks Joe,
I subscribed to your blog just based on this post. Excellent criticisms. I hope you do end up submitting a letter to Science.
rory, Tuuli,
Thanks for the comments; a comment to Science is indeed in the works.
Hi there. I like your comments on this topic. Sounds fairly reasonable in general. We are a proteomics lab, so I tend to come at this from a slightly different perspective. My line of thinking is that unless you show RNA edits are translated to proteins, then you really can’t be sure of anything. Li et al go some way to addressing this point by doing gelC-MS/MS shotgun proteomics on pooled lysates from B cells derived from 15 CEPH patients. However, calculating peptide false discovery rates (FDR) from this type of dataset is fraught with difficulties and the jury is still out on the best way to do this.
Given the uncertainty surrounding false positives in shotgun proteomics, I would love to see them do selected reaction monitoring (SRM/MRM)(using a triple quad mass spec) on a set of “mismatch” peptides, using stable isotope labeled versions as standards. This is truly the gold standard for quantitative evaluation of peptide abundance and it would allow us to compare relative amounts of “normal” versus “edited” peptide. With the proteomics data provided in the manuscript we really can only say if a given peptide was detected or not – i.e. no quantitation. Also they don’t show any MS/MS spectra for edited peptides – this would be nice to look at to see how “good” they look.
In summary unless they can validate their shotgun results by doing targeted quantitative proteomics on at least a subset of mismatch peptides, then a large question mark remains.
Hi Patrick,
Thanks for the comment.
Also they don’t show any MS/MS spectra for edited peptides – this would be nice to look at to see how “good” they look.
Agreed. The fact that the primary MS data is not present in the paper is something even I (someone with no experience with proteomics data) found a bit odd.
Just a comment on their selection criterion of the editing event.
It is important to point out than their criterion for defining an editing event is rather weak, at least 10% of the reads from RNA-Seq needs to carry the base that is different from the DNA sequence. They were considering all exonic positions that are covered with at least 10 RNA-Seq reads, which theoretically leaves a possibility they were analyzing positions with only one RNA-Seq read which contain a mismatched base (10% of 10 reads). This indicates they might have been analyzing a significant number of sequencing errors, if I correctly understood their explanation.
Hi MP,
To clarify, in order to report an RDD site, they required that it meet their selection criteria in at least two individuals. This means they required a minimum of 2 reads covering the site. After reprocessing their raw data, I can tell you that ~7% of their RDD sites have 2 mismatching reads, and ~23% have less than 5.
Hello, just wondering if there is an update for this exciting story, particularly in light of below recent paper?
Extensive genomic and transcriptional diversity identified through massively parallel DNA and RNA sequencing of eighteen Korean individuals – pp745 – 752
Young Seok Ju, Jong-Il Kim, Sheehyun Kim, Dongwan Hong, Hansoo Park, Jong-Yeon Shin, Seungbok Lee, Won-Chul Lee, Sujung Kim, Saet-Byeol Yu, Sung-Soo Park, Seung-Hyun Seo, Ji-Young Yun, Hyun-Jin Kim, Dong-Sung Lee, Maryam Yavartanoo, Hyunseok Peter Kang, Omer Gokcumen, Diddahally R Govindaraju, Jung Hee Jung, Hyonyong Chong, Kap-Seok Yang, Hyungtae Kim, Charles Lee & Jeong-Sun Seo
doi:10.1038/ng.872
Jeong Sun-Seo and colleagues report whole-genome sequencing of ten Korean individuals and exome sequencing on an additional eight Korean individuals. They also performed transcriptome sequencing on 17 of these individuals. The authors identified approximately 1.83 million previously unidentified SNPs.