The first thing I did when I received my genotyping results from 23andMe was log on to their website and take a look at my estimated disease risks. For most people, these estimates are one of the primary reasons for buying a direct to consumer (DTC) genetics kit. But how accurate are these disease risk estimates? How robust is the information that goes into calculating them? In a previous post I focused on how odds ratios (the ratio of the odds of disease if allele A is carried as opposed to allele B) can vary across different populations, environments and age groups and, as a consequence, affect disease risk estimates. It turns out that even if we forget about these concerns for a moment, getting an accurate estimate of disease risk is far from straightforward. One of the primary challenges is deciding which disease loci to include in the risk prediction and in this post I will investigate the effect this decision can have on risk estimates.
To help me in my quest, I will use ulcerative colitis (UC) as an example throughout the post, estimating Genomes Unzipped members’ risk for the disease as I go. Ulcerative colitis is one of two common forms of autoimmune infllammatory bowel disease and I have selected it not on the basis of any special properties (either genetic or biological) but because I am familiar with the genetics of the disease having worked on it extensively.
The table below gives our ulcerative colitis risks according to 23andMe. The numbers in the table represent the percentage of people 23andMe would expect to suffer from UC given our genotype data (after taking our sex and ethnicity into account). The colours highlight individuals who fall into 23andMe’s “increased risk” (red) or “decreased risk” (blue) categories based on comparisons with the average risk (males: 0.77%; females 0.51%). As far as I am aware none of us actually do suffer from UC.
| Carl | Caroline | Don | Dan | Ilana | Jeff | Joe | Jan | Kate | Luke | Vincent | Daniel |
| 0.72% | 0.44% | 1.13% | 0.67% | 0.44% | 1.09% | 1.43% | 0.55% | 0.44% | 0.89% | 0.66% | 0.31% |
One of the more difficult decisions that DTC companies are faced with is deciding which loci to include in their risk models. As someone who has spent a lot of time trying to identify loci associated with UC, I was a little bit disappointed to find out that 23andMe only include four loci in their risk model. There are currently 47 confirmed UC loci and the table below gives our UC risks if all of these are included in the prediction algortihm.
| Carl | Caroline | Don | Dan | Ilana | Jeff | Joe | Jan | Kate | Luke | Vincent | Daniel |
| 0.48% | 0.51% | 0.61% | 1.30% | 0.61% | 1.14% | 0.34% | 0.43% | 0.14% | 0.40% | 0.65% | 0.11% |
When comparing these results to those in the previous table the first thing to note is that for some of us the risk prediction does not change a great deal (Caroline still has a delightfully uninteresting genome). For others, using all 47 confirmed UC loci in the prediction has changed things substantially. When Joe logs on to the 23andMe website he finds UC in the ‘Elevated risk’ list of diseases, but my analysis shows that (when using all available markers) it should actually be listed under ‘Decreased risk’. Joe’s 23andMe prediction is heavily influenced by the fact that he is homozygous for the risk allele at BSN (one of the four loci included in the 23andMe prediction) but when all 47 loci are considered the influence of this one locus dissipates (the same is true for Don). For others the news is not so good. Dan previously thought that he had an average risk of UC but my analysis shows that his risk is actually 1.69 times above the average (though his absolute risk is actually still low, so I don’t imagine he will be losing any sleep over this).
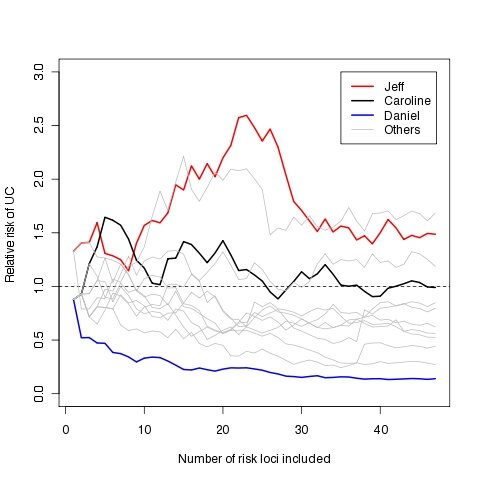
The graph below shows how our relative risks of UC change as the number of risk loci included in the predictive algorithm is increased. The loci are added in such a way that the most important in terms of UC risk prediction gets added first and then so on until all 47 are included. I have higlighted the UC relative risks for Jeff, Caroline and Daniel as examples of elevated, typical and decreased risks, respectively. The rest of us are shown in gray. As you can see, in some cases relative risk can vary quite substantially depending on the number of loci included in the risk model, but broadly speaking we seem to be well classified as increased, decreased or typical risk using only 5-10 of the most predictive loci (this number will vary between traits).
So why does 23andMe only include four markers in their UC risk prediction algorithm? In their whitepaper ‘Vetting Genetic Associations (June 2010)‘ 23andMe state that to be included in their prediction algorithms loci must be replicated in ‘at least one independent published study’. Before the advent of genome-wide association studies (GWAS) this was certainly a necessary step because candidate-gene studies were notorious for turning up false-positive findings that were difficult to replicate. The statistical rigour that has accompanied GWAS has reduced the number of false-positive findings and successful replication must be demonstrated before a GWAS can be published in top-tier journals such as Nature Genetics or New England Journal of Medicine. But there is the crux, for the majority of loci being robustly identified via large-scale meta analysis there will never be an independently published replication study (the replication study will be published together with the ‘discovery’ GWAS meta-analysis). The loci being highlighted by these studies can take tens of thousands of samples to identify and there is simply not another cohort of this size lying around waiting to take part in an independent replication study. I would advise 23andMe to remove the need for independently published replication studies. Providing the replication study uses independent samples and a different genotyping technology then I have no issue with these being reported in the same manuscript as the discovery cohort. If this were adopted, 23andMe risk predictions would include the vast majority of loci identified by meta-analyses and provide us all with the best genetic estimate of our disease risk possible at this time.
In part two of this post (available soon) I will focus on other, perhaps more technical, factors involved in risk modelling and investigate how robust our disease risk estimates are to small perturbations in these.








 RSS
RSS Twitter
Twitter
Been waiting for something like this to be published. Credibility of 23andMe in the physician community would have improved if they had published a report like this in (their blog) The Spittoon.
Great post, Carl.
One interesting idea that DTC companies could consider: allow customers to set their own evidence thresholds. Just have a little toggle that you can switch from super-conservative (e.g. only independently replicated variants) to very relaxed (as an extreme, everything that’s popped up with a P value below 10^-5, for instance) and various points in between (for instance, allow for SNPs from meta-analysis). Then customers could get a sense of how much these inclusion criteria affect their risk predictions, and decide for themselves what threshold they think is most reasonable.
Don’t think companies like 23andMe haven’t already thought of all of these things. It’s not usually so simple as “just put in a toggle”. There are many other issues – regulatory, engineering, and UI – to consider on top of the scientific/policy considerations that interest this crowd, that make it actually a pretty complicated feature to implement. You also have to remember that the average user may not know right away how to use or make sense of such a feature, or may not want it. That isn’t an argument to not do it, it just means the implementation needs to be thoughtful and accommodating to more than just one type of user. (Some might say that’s an argument FOR the toggle — so that people who want different levels of evidence can get it — but the people who know what kind of evidence they want make up one very particular type of user.)
That said, these are all reasonable suggestions. :)
Hi Shirley,
Sorry, I didn’t mean that to sound so flippant – “just have a little toggle” obviously elides a huge number of challenges, as you say. And of course the geeky features I’d like to pack into the UI (if I had, say, infinite engineers at my disposal) may well be of interest to only a small fraction of other customers.
So that’s very much an optional extra, cool as I think it would be. Coming up with a way of cleanly adding in variants from large meta-analyses seems rather more urgent.
One problem with allowing different levels of confidence it seems is that there is a bias-variance tradeoff: it should be the case that the odds ratios for SNPs with larger p-values will tend to be more inflated over their “true” value than those who comfortably clear the threshold. You can view 23andMe’s white paper as essentially an effort to remove as much bias as possible from the results (not saying that’s the right thing to do, but…)
Globally there are tradeoffs like this too: one of the nice parts about requiring replications is that you have a chance to see the SNP typed twice, so you have some confidence that the authors got the rsid, risk allele, and odds ratios all correct. Would you give up some power to be more sure about the results?
Whether or not this is the correct approach is difficult. Certainly here there won’t be a replication, and 4 vs 47 SNPs is a compelling difference. But how much does it raise your prediction accuracy? Or substitute “utility” for “prediction accuracy” if you want a harder problem :)
Interesting topic, as always. I love this blog.
Regarding the current topic: we have written a paper on how risk change when predictions are updated by adding more variants.
See: Mihaescu R, van Hoek M, Sijbrands EJ, Uitterlinden AG, Witteman JC, Hofman A, van Duijn CM, Janssens AC. Evaluation of risk prediction updates from commercial genome-wide scans. Genet Med. 2009 Aug;11(8):588-94.
Great post. What prediction algorithm have you used for the inclusion of other SNPs? Are there any script available?