In my last post, I discussed how I used 23andMe data to test hypotheses about my ancestry. In particular, I was intrigued by Dienekes Pontikos’s result suggesting that I (and my colleague Vincent) might be partly Ashkenazi Jewish. Ultimately, however, I concluded that his algorithm was not properly modeling my southern European ancestry (inherited from one Italian grandparent), and that this was leading to a spurious result.
I was wrong.
What did I conclude previously?
Let’s quickly recap my previous discussion: Dienekes’s program models individuals as being a mixture of Ashkenazi Jewish, northwest European, and southeast European ancestry. People, like Vincent and myself, who are not fully descended from those three populations will pose problems for this algorithm. I thought it was unlikely to be a coincidence that Vincent and I were the only two people to get confusing results. Indeed, when I included Italian and French individuals in my analysis, I saw no clear evidence for any Ashkenazi ancestry on either of our parts. Mystery solved.
Some inconsistencies revealed
After I published that previous post, however, a couple things came up that seemed incongruous. First, a commenter recommended that I check out the Ancestry Finder tool on 23andMe. What this tool does is identify large segments of your genome that perfectly match the genomes of other people of known ancestry. If, for example, parts of my genome perfectly matched an individual who knows for a fact that s/he is Ashkenazi, this would be pretty strong evidence that those parts of my genome were descended from someone who also was Ashkenazi. Indeed, this is what I found—a moderate proportion (3-30%) of my genome does indeed appear to be of recent Ashkenazi ancestry in this analysis. I was skeptical about this, but on reflection, I couldn’t come up with a good reason that this result would be spurious.
Second, Dienekes followed up on his analysis of the ancestry of the GNZ participants with a much larger data set, including individuals of southwest European descent. As expected, when including more data, there was no evidence that Vincent has any Ashkenazi ancestry. Unexpectedly, this was not true for me—even in this larger analysis, the evidence for Ashkenazi ancestry didn’t disappear.
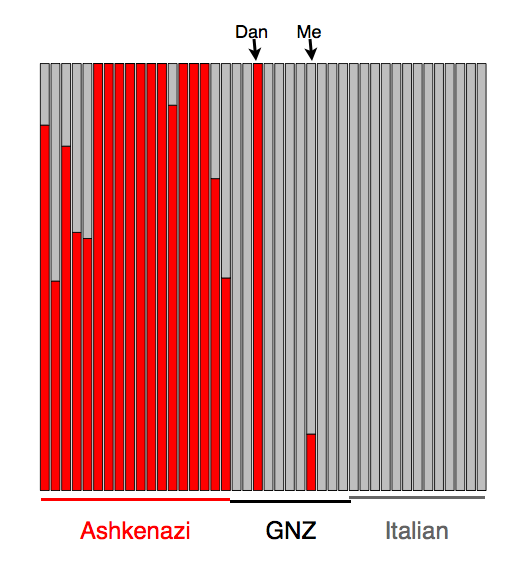
I followed up on this using a similar approach to Dienekes. I used the same dataset I assembled previously—a set of European populations from the Human Genome Diversity Panel, a set of Ashkenazi individuals, and the GNZ data. This time, instead of using principle components analysis (which averages information across the entire genome), I used the model implemented in the program admixture (which models individuals as mixtures of different populations) [1]. With this model and these data, it’s relatively easy to find a component of ancestry that is essentially unique to the Ashkenazi population [2]. Below, I’ve plotted (in red) the estimated fraction of Ashkenazi ancestry for a subset of individuals from this analysis. As you can see, there are two GNZ individuals with any red: Dan (who knows he is fully of Ashkenazi descent) and, surprisingly, myself. Combined with the Ancestry Finder results, there are two possibilities: either all the algorithms are getting confused (one can imagine situations where this would be the case) or I’m confused myself.
The resolution
As I was mulling over these sorts of issues, I sent the link to my previous analysis to a family member. I didn’t really expect this person to find it that interesting, but hey, you never know. I then got a phone call. I’ll summarize a couple days worth of moderate confusion, second-hand reports of conversations with distant relatives, and family intrigue with this: as it turns out, one of my great-grandparents was indeed a Polish Ashkenazi Jew who immigrated to the United States around the turn of the century. I, obviously, was completely unaware of this.
So to conclude, a tip of my hat to Dienekes and everyone else who looked at these data—this has been the first genuinely unexpected thing to come out of my genetic data.
—
[1] Alexander et al. (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Research. doi: 10.1101/gr.094052.109
[2] I first thinned the data to remove SNPs in strong linkage disequilibrium. I then ran admixture using K=8, 9, and 10, looking for an ancestry component essentially specific to the Ashkenazi population. The program finds one at K=9. Plotted in red in the figure is the fraction of each individual’s ancestry predicted to be from this population (which I’m interpreting as Ashkenazi). I ran this on all the individuals, but am only plotting the GNZ individuals, and the Ashkenazi and Italian populations for comparison.








 RSS
RSS Twitter
Twitter
wow. that’s pretty awesome.
probably unlikely, but i wonder if this sort of thing could be integrated into the law of return:
http://en.wikipedia.org/wiki/Law_of_Return
Agree with Razib, that is pretty awesome. Have you already checked for Tay-Sachs?
I can’t believe I’m asking such a personal question, but it seems on-topic to ask.
Razib,
The Law of Return is only with a full Jewish grandparent. This was Hitler’s criteria for extermination. My guess is that someone who brought genetic evidence would be accepted no problemo.
However, they would not get stamped “Jewish” on their ID card unless the Orthodox approved, and even if one brought evidence that their mother’s mother was Jewish he would not be accepted unless they themselves were at least moderately observant.
Have you already checked for Tay-Sachs?
According to 23andMe, I’m not a carrier. I haven’t looked at it carefully, but I imagine they’re looking for the variant that’s common among the Ashkenazi.
Again, thanks for sharing this. Now you’ve got your Oprah moment!
So *this* is what you do in your free time, eh? UChicago represent! :)
Very cool results, though. In theory you should be looking at other mutations besides Tay-Sachs, right? BRCA mutations, for example.
Hey Jessica. Yes, in my time off from doing genetics, I…do genetics :)
I’m not a carrier for anything tested by 23andMe, though their test certainly isn’t exhaustive.
Out of curiosity about the non-science/social aspects of what you’re doing:
Did you feel uncomfortable when I asked about Tay-Sachs? At first, I was thinking about the data much like I do about any data: I wonder what interesting things one could learn from it. So I asked what I thought was a natural question to ask. But after I wrote that question, I was like, “what?! I’m asking someone about an incredibly personal issue.”
If you were a carrier for anything tested by 23andMe, would you feel comfortable talking about it? Some of the other genomesunzipped participants have talked about the lines that they’ve personally drawn around their data, but I don’t recall reading your take.
Or more generally: I don’t think I know what the etiquette/protocol should be w.r.t. what’s in-bounds or out-of-bounds for this discussion.
Mitch,
Feel free to ask me anything about my data. I’d have no problem saying I’m a carrier of any disease alleles; I presumably have a few that aren’t being testing by 23andMe.
@Mitch: You wrote “Some of the other genomesunzipped participants have talked about the lines that they’ve personally drawn around their data, but I don’t recall reading your take.”
I’m curious what you mean.
@Joe: Nice to know I’m not the only one with Ashkenazi heritage in the group, even if it took some digging to confirm it!
even if it took some digging to confirm it!
wait. are you implying that joe tweaked your “jewdar”? or is joe’s connection to the tribe as surprising to you as it was to the rest of us goyim? :-)
Joe,
thanks for the post, it’s really great. Can you then figure out what parts of your chromosomes were inherited from your Ashkenazi great grand-parent? And make a nice chromosome painting type plot? The blocks should still be large enough to identify them well and it would be a nice thing to show.
@Dan:
I’m curious what you mean.
I didn’t phrase that very well, but I was thinking about your post, where you talked about how you felt that it was necessary to get consent from your family members to have your data here. I also had a vague idea that someone here had asked that something about their data not be discussed, but now I realize that I was actually thinking about how Jim Watson didn’t want to know his APOE status.
Generally speaking, I wasn’t sure what the boundaries (if any) are. Is everything open to discussion here, or do any of the genomesunzipped participants prefer to avoid certain topics? If, say (to give a contrived example) I read a new paper that links a certain genetic variant with erectile dysfunction, and I find that one of you has that variant, is it okay to ask how often you experience ED? Is it okay to write a blog post about it, with your name attached?
I almost certainly wouldn’t do that, of course, because it strikes me as being pretty rude, but I don’t know quite where the line is.
Joe,
It was clear to me you have Ashkenazi ancestry when I looked at your HIR Search results on http://hirs.snpology.com/
(You can read about the HIR Search in Friday Links https://genomesunzipped.org/2010/10/friday-links-8.php )
Both “Show Relatives” and “Detailed Reports” options show a solid number of Ashkenazi cousins. No doubts at all…
Ha, Mitch, that would indeed be a bizarre and uncomfortable situation.
I say we cross that bridge when we come to it–in general, I’ve placed the probability of actually finding anything awkward or upsetting in my genome at about 0; if I feel differently at some point we’ll see.
Vincent,
That’s a good idea, and along the lines of what Ancestry Finder at 23andMe does. Doing it myself would require quite a bit of work, which I’m unlikely to do anytime soon. :)
All this is fascinating indeed. And the mention of the law of return clearly shows the potential socio-ethical issues generated by such tests. Another example is “Native American testing” (http://www.sciencedaily.com/releases/2007/10/071018145955.htm) and its potential impacts (from financial benefits to serious identity issues…)
I was wondering if you and your “unzipped” ;-) colleagues had any concern about (unwillingly) promoting direct-to-consumer testing (and thus private firms services…), given the current and growing controversies it raises.
Best,
Thierry
The Law of Return is only with a full Jewish grandparent. This was Hitler’s criteria for extermination.
True.
My guess is that someone who brought genetic evidence would be accepted no problemo.
False. Genetic test results indicating plausible Jewish descent on the Y chromosome won’t cut it with Israel’s Interior Ministry to prove Jewish ancestry for purposes of immigration. Additionally, you have to show that it is not longer than 3 generations away. You can qualify for aliyah under the Law of Return if you can show convincing documentary evidence (community records, Jewish marriage contract, school records, burial records) that you are descended from at least one grandparent who was a member of a recognized Jewish community somewhere.
Perhaps I wasn’t clear. I suspect that if someone brought genetic evidence that they had a FULL Jewish grandparent, that they would be accepted as Israeli citizens (but not stamped as Jews acc to Jewish law). I did not mean to suggest that someone who brought evidence of previous Jewish ancestry, such as in Joe’s case here, would be accepted, which Razib asked about.
Generally speaking, I wasn’t sure what the boundaries (if any) are. Is everything open to discussion here, or do any of the genomesunzipped participants prefer to avoid certain topics?
No questions are off-limits, but of course members can choose not to answer them. :-)
If, say (to give a contrived example) I read a new paper that links a certain genetic variant with erectile dysfunction, and I find that one of you has that variant, is it okay to ask how often you experience ED? Is it okay to write a blog post about it, with your name attached?
If it’s something you’d feel uncomfortable having shared about yourself, you can always email us to have a chat about it first – I doubt any of us would ask you not to write about it, but we’d appreciate the advance warning. One way of softening the blow would be to refer to the individual by their GNZ ID number (e.g. JCB001) in your post rather than their name, so at least someone Googling Jeff doesn’t find “Jeff Barrett carries a genetic variant for erectile dysfunction” on the first page.
But there aren’t any specific rules here, and we all understand that there will be people who don’t ask permission before disclosing uncomfortable facts about us – that’s just a risk of the project.
I’m ambiguously brown and your earlier post on this really resonated with me. I think that your analytical journey on this one from start to finish is really informative, and the end result is cool! I’m jealous.
As you say, this unexpected new knowledge did come out of your genomic data, BUT it was also the product of your know-how (which allowed you to start corroborating Dieneke’s analysis) and it was your relationship with a relative that seemed to seal the deal for you. I guess my question is where you imagine you would be in your thinking if you didn’t talk to your family much and had stopped at the nice clear graph?
This is probably a thought experiment I should keep to myself, but I do wonder what it will be like when neonazis get into ancestry testing, I bet it will really mess them up.
Is it too late for you to wrangle your way into Birthright?
I guess my question is where you imagine you would be in your thinking if you didn’t talk to your family much and had stopped at the nice clear graph?
That’s an interesting question; without talking to my family I’m not sure how I would have interpreted this data. My guess is that I would have remained skeptical, but I’m not sure. I had no previous knowledge of Jewish ancestry, so it really took overwhelming evidence to convince myself otherwise. If you have some suspicions about your ancestry, maybe genetic evidence alone would convince you. So I’m not sure. Sorry to be so vague :)
This is probably a thought experiment I should keep to myself, but I do wonder what it will be like when neonazis get into ancestry testing, I bet it will really mess them up.
oddly enough, Razib pointed to a story along these lines just the other day:
http://www.aolnews.com/world/article/neo-nazi-couple-find-out-theyre-jewish/19648414
I know an “Ashkenazi” Jew, whose YDNA matches show very strong evidence for a Spanish ancestor who probably fled the inquisition. This has been very meaningful to him.
http://truthinjustice.ning.com/forum/topics/many-ashkenazi-jews-have?xg_source=activity
Also, someone who tested mtDNA and found matches “like a synagogue seating list”, investigated, and confirmed Jewish matrilineal lineage. This person now feels affiliated as a Jew, but struggling with issues of dogma.
Also, those who thought they had a pedigree, turns out they don’t, and are bummed out.
Razib wrote a good post on the importance of this info for purposes of identity. Like it or not, this is who you were before you were you.
“One way of softening the blow would be to refer to the individual by their GNZ ID number (e.g. JCB001) in your post rather than their name, so at least someone Googling Jeff doesn’t find “Jeff Barrett carries a genetic variant for erectile dysfunction” on the first page.”
Daniel, I guess you realize that you just wrote down that sentence and Google sees these discussions too :-)
Daniel, I guess you realize that you just wrote down that sentence and Google sees these discussions too :-)
Why, so I did!
Yes, I’m a bad, bad person…
Joe, which of the products of 23andMe did you, and the other GNZ participants, use for your ancestry data? I recently recieved results of a 27 STR autosomal marker test, from DNA Tribes, and was astounded to see an affiliation with Bedouins, (Negev, Israel) running at 20% of the strongest affiliation, which was Iceland. This was on both their “Native” and “Global” population matches. On their “World Region” match results I was even more amazed to have the strongest match be “Arabian”, followed closely by “Eastern European”. These results are baffling as all my ancestors are from the British Isles and Germany.
I would like to see how this 27 marker test stacks up with 23andMe’s test, which, I think, uses hundreds of thousands of autosomal SNPs.
@David
All of the GNZ contributors got the 23andMe Complete package, which gives raw data on 560k autosomal SNPs (plus ~4000 Y and MT SNPs).
We don’t have any DNA Tribes data for our group, so I have no idea how accurate it is. It feels like using only 27 STRs would raise a small but significant probability of getting false positive ancestry results.
David, take into account that you can order the FTDNA FF test (548k SNPs) which is cheaper than 23andme one. If you’re not interested in Y and MT SNPs, it may worth to take it.
Luke, Leon: Thanks for the info.
David: The biggest problem with the dnatribes results is that they don’t work very well for admixed people as they are trying to match you to a single population. I get consistent overall results with dnatribes, BGA analysis of my 23andMe and Family Finder data, and EHSTRAFD (where you can input your STR markers to get a free second opinion on your results). The challenge is to unpack one’s overall data into the different global populations that you descend from (detailed admixture analysis which is at best experimental right now).
Helen: Thankyou for the observations. But what do the acronyms BGA and EHSTRAFD stand for?
Last year I had my Y-DNA tested, which turned out to be E1b1b1, with three of the four strongest matches tracing their roots to the Catalonian region of Spain, at 36/37, 25/25, 24/25. That made me think that my direct paternal ancestor came from that part of Spain in the distant past, even though my dad’s ancestors resided in a small German village for the last 300 years, where the paper trail ends about 1705.
In case anyone has the time, or inclination, to re-analyze my DNA Tribes results, here they are:
Locus Allele 1 Allele2
Amel X Y
D3S1358 15 18
TH01 9 10
D21S11 28 30
D18S51 12 17
Penta E 9 10
D5S818 11 11
D13S317 11 13
D7S820 9 11
D16S539 11 12
CSF1PO 10 10
Penta D 9 11
vWA 17 17
D8S1179 12 12
TPOX 8 11
FGA 23 24
D2S1338 17 25
D19S433 15 16
F13A1 7 7
F13B 9 10
FES/FPS 10 13
LPL 10 11
D10S1248 14 17
D12S391 18 24
D1S1656 11 17
D22S1045 15 15
D2S441 11 13
SE33 26.2 26.2
I typed this data in by hand, so am not sure if it will show up as a neat list.
BGA = biogeographical ancestry, clues inferred about the geography of our ancestors by comparing our autosomal DNA with that of reference population samples from different geographical areas
EHSTRAFD = Earth Human Short Tandem Repeat Alleles Frequency Database (Most Probably Geographic Origin), submit your STR markers here for analysis: http://www.ehstrafd.org/modules/mpgo/form.cfm
However, like dnatribes it will only try and match you with one single population at a time which is problematic if you have mixed ancestry.
Your YDNA is only one out of many ancestral lines, don’t place too much emphasis on it for BGA purposes.
You can also try asking for advice in DNA Forums (dna-forums.com) in the Autosomal DNA section.
That should read Most Probable Geographic Origin, I don’t know why but I always type it wrong.
Helen: Thankyou so much for referring me to that website. I just inputted my data, and got the following, if you don’t mind reading a laundry list: 1) Polish (Northeast Poland) 100, 2) Polish (Central Poland) 64.37, 3) Venezuelan (Maracaibo) 59.03, 4) Belgium 43.37, 5) Saudi Arabian (Dubai Emirate) 42.41, 6) Polish (Lodz) 38.24, 7) Polish (North-Central Poland) 28.14, 8) Caucasian (United States) 26.79, and so on. Poland even comes up on number 10.
You were absolutely right not to take to much stock in the Y-chromosone results. It looks like Poland is the top billing, which really isn’t too surprising, since Germany borders Poland, and no doubt there’s been a lot of population movement between the two countries. But, it’s interesting that a Hispanic country is in number 3 position, suggesting a strong input from that group of people. This analysis was run with the “All” function selected. I haven’t tried the “Mixed”, or “Region” functions yet. Thanks again.
Helen: I just tried the “Mixed” population function and the results changed somewhat. In order: Polish, Polish, Venezuelan, Belgium, Polish, Polish, Croatian, Polish, Mestizo. Saudi Arabia disappeared from the list altogether. I misnamed the other population function, it was “Labeled”, not “Region”. Maybe this is a more accurate picture, considering I have mixed ancestry from the British Isles and Germany.
David: Your EHSTRAFD laundry list is very similar to mine and my ancestry is also British and Germanic (whew, that’s a relief). I think it would probably be best to continue this discussion in DNA Forums so as not to hijack Joe’s post?
BTW there is another similar free STR matching program I forgot to mention i.e. Omnipop (See en.wikipedia.org/wiki/OmniPop and http://www.cstl.nist.gov/biotech/strbase/populationdata.htm).
At this stage 23andMe and FTDNA only provide admixture analysis at a fairly high level and the genetic genealogy community is pushing for greater detail (I am 100% Euro according to both and predominantly Western Euro according to FTDNA). Fortunately for us various independent “academics” such as Dienekes are pushing the envelope.
DNA Tribes provides detail but there is a huge trade-off against accuracy. Unless all your ancestors come from the same place, take these results with a big dose of salt.
Oops forgot this one too: strbase.org/calc.php
P.S. In my experience unless you have known ancestry from there Spain, Basque and Hispanic results are all basically indicators of deep Euro ancestry and/or a typical Western Euro genetic profile. I am British-Germanic and I consistently tend to average out in Spain/France/Belgium.
Helen: Thanks for all the new information. I’ll check out those other two STR matching programs. Last nite I realized I was getting off-topic for Joe’s post, so I’ll head on over to DNA-Forums.
Very good information. Lucky me I ran across your blog by accident (stumbleupon).
I’ve book-marked it for later!